Full documentation for this project can be found on the Github Pages site
Introduction
During my PhD, one major project I had was to create a statistical shape model of the lungs. This involved running a multi-step workflow using a large dataset of lung scans.
The software I wrote for this project (written in Python) is composed of both code for creating the shape model itself and some tools I developed to help orchestrate my workflows. The code is hosted in a public Github repository, and documentation generated with Sphinx is available on a Github Pages site.
Dataset Configuration
The workflow orchestration code is designed to work with a specific type of dataset called the SPARC data structure, which consists of nested directories along with associated metadata files.
dataset_name/
|--- dataset_config.json
|--- primary
| |--- subject_directory_1
| |--- subject_data.nii
|--- derivative
| |--- subject_directory_1
| |--- raw_mesh
| |--- lobe_1.stlThe block above illustrates the dataset format. There is a root directory with a nested structure that contains directories for each human subject. Initial data is stored in the primary directory, and a mirrored directory structure is created to associate any derived data with its original subject.
To help my workflow orchestrator parse the dataset, I introduced a dataset configuration file as seen below. This indicates to the program which folders to identify as primary and derivative, how deep the nested structure is, and what to use for subject identification.
{
"primary_directory": "primary",
"derivative_directory": "derivative",
"pooled_primary_directory": "pooled_primary",
"pooled_derivative_directory": "pooled_derivative",
"directory_index_glob": "directory_index*.csv",
"data_folder_depth": 2,
"subject_id_folder_depth": 2,
"lung_image_glob": "*.nii",
"lobe_mapping": {"rul": 3, "rml": 4, "rll": 5, "lul": 1,
"lll": 2}
}Workflow Configuration
The core of the application is the workflow, and in this project each new workflow is defined using a separate workflow configuration file, which contains all the instructions needed to run it. The workflow configuration file consists of a series of tasks listed by a name corresponding to code in the repository (which I will cover shortly). The tasks communicate with each other by reading and writing files in each task’s configured source and results directory. The configuration file also contains any user-configurable parameters available to adjust the behavior of the tasks. Below is an abbreviated example of a workflow configuration file:
dataset_root: null
log_level: "INFO"
use_multiprocessing: False
tasks:
initialize:
dataset_config_filename: "dataset_config.json"
use_directory_index: False
skip_dirs: [ ]
select_dirs: [ ]
extract_whole_lungs_sw:
task: "ExtractWholeLungsSW"
results_directory: "extract_whole_lungs_sw"
output_filenames: { left_lung: [ "lul", "lll" ], right_lung: [ "rul", "rml", "rll" ] }
params: { maximumRMSError: 0.009999999776482582, numberOfIterations: 30 }
create_meshes_sw:
task: "CreateMeshesSW"
source_directory: "extract_whole_lungs_sw"
results_directory: "create_meshes_whole_lungs_sw"
image_glob: "*.nii"
params: {
pad: True,
step_size: 1,
decimate: True,
decimate_target_faces: 100000,
subdivide_passes: 0,
volume_preservation: True,
remesh: True,
remesh_target_points: 40000,
adaptivity: 0,
smooth: True,
smooth_iterations: 10,
relaxation: 1,
fill_holes: True,
hole_size: 100,
remove_shared_faces: True,
isolate_mesh: True }
logging:
run_tasks: [ "extract_whole_lungs_sw", "create_meshes_sw"]Software Process
Workflows are run using a command line application included in the repository. The first step of any workflow is the initialization process (shown below) The configuration file provides a list or glob of subject ids that the workflow should act on, and the application searches through the dataset to find a list of absolute directories to reference.
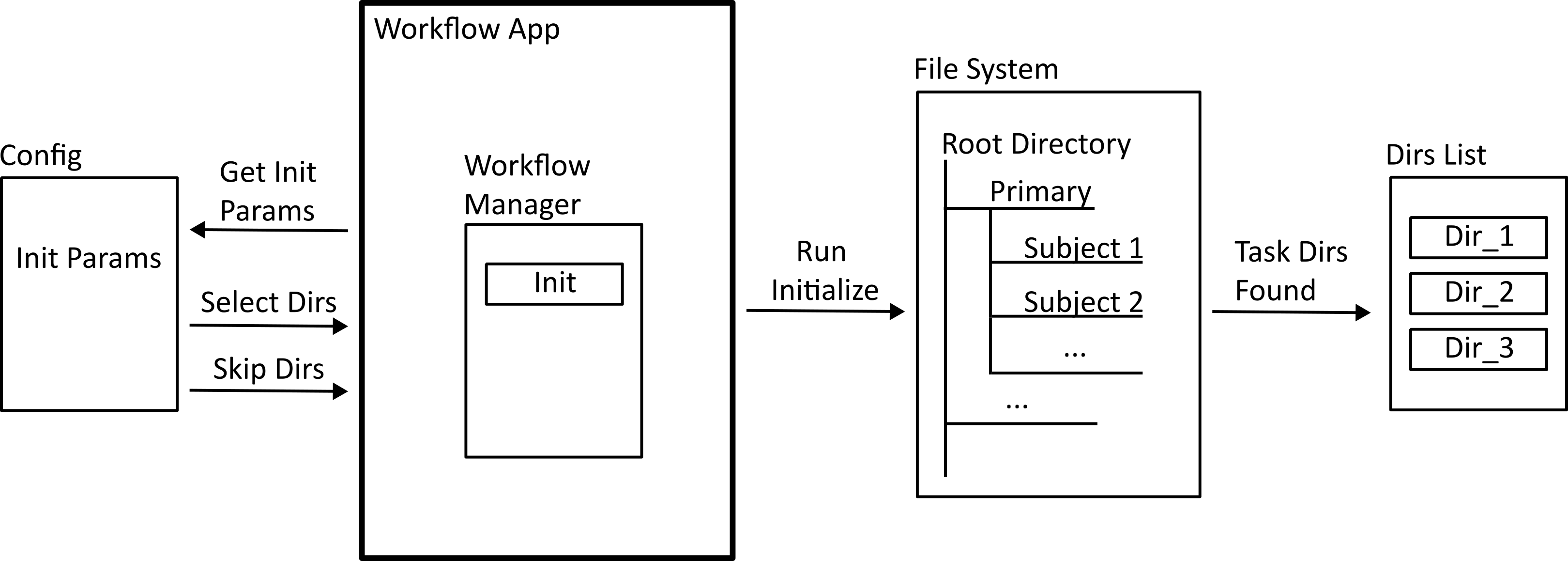
The next step is task registration. The configuration file provides the list of tasks in the workflow by name, and the application needs to find the code that these represent. The code for each task needs to implement a standard interface so that they can be handled properly by the workflow manager. In Python, this is done by defining an abstract class which each task needs to implement.
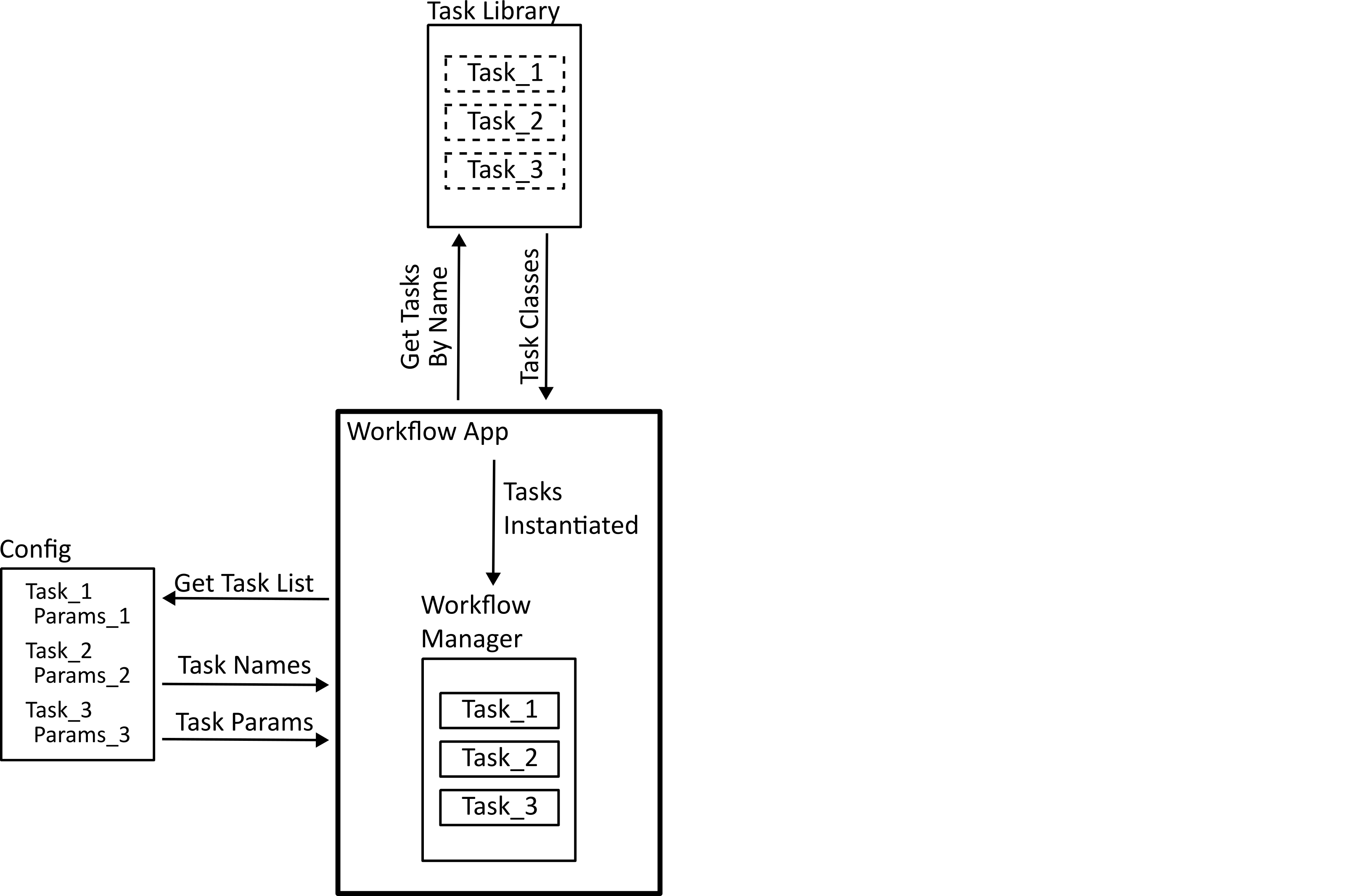
Finally, the workflow is run. Tasks that run on a single subject individually can be dispatched to a multiprocessing mapper to run in parallel. Before that happens, the function is wrapped using an exception monitoring decorator so that if there is an error in the function it can be reported back to the main process.
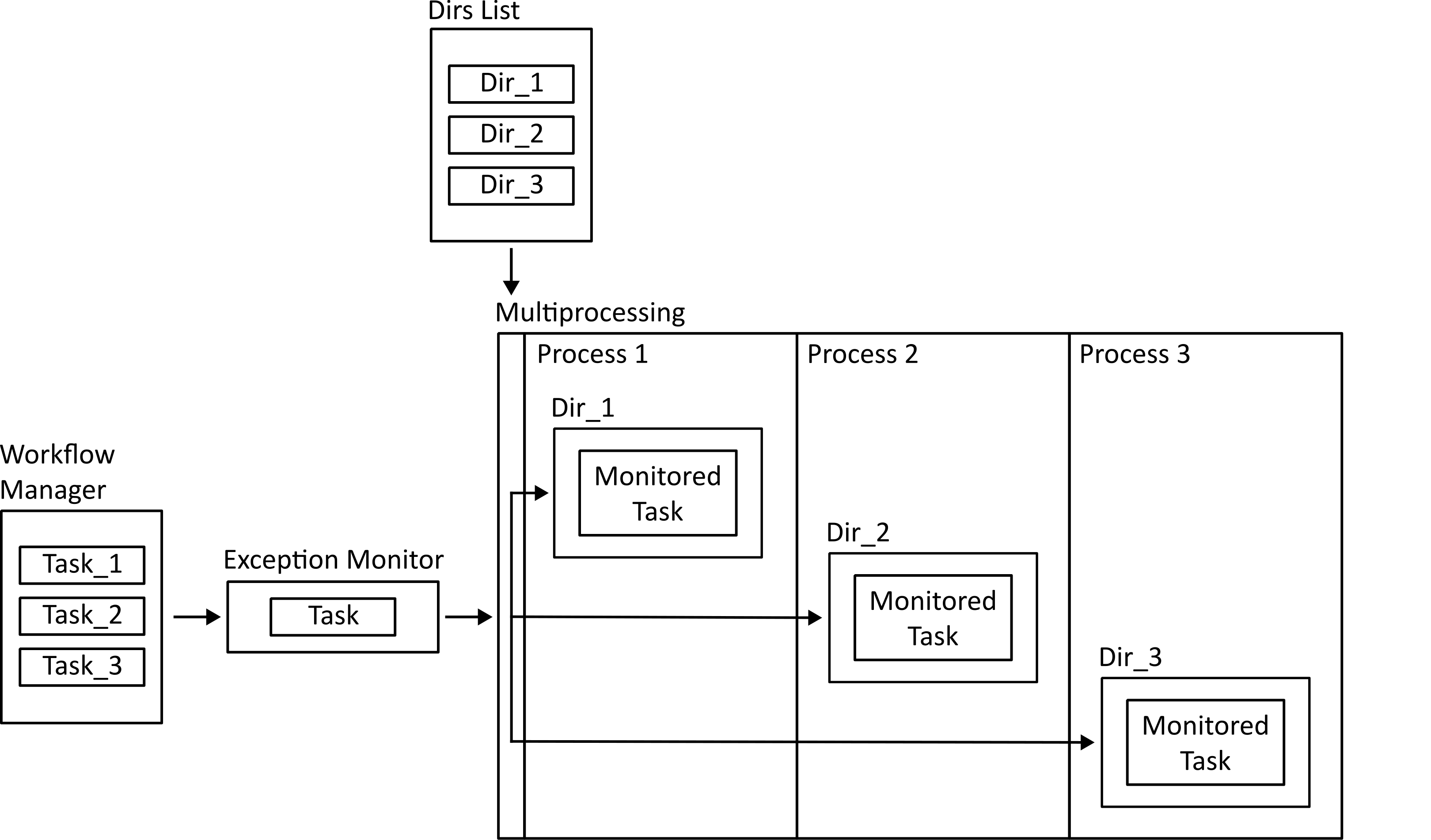
Tasks that need to run on the ensemble of subjects are run in the main process. They are still monitored for exceptions so that errors can be logged.
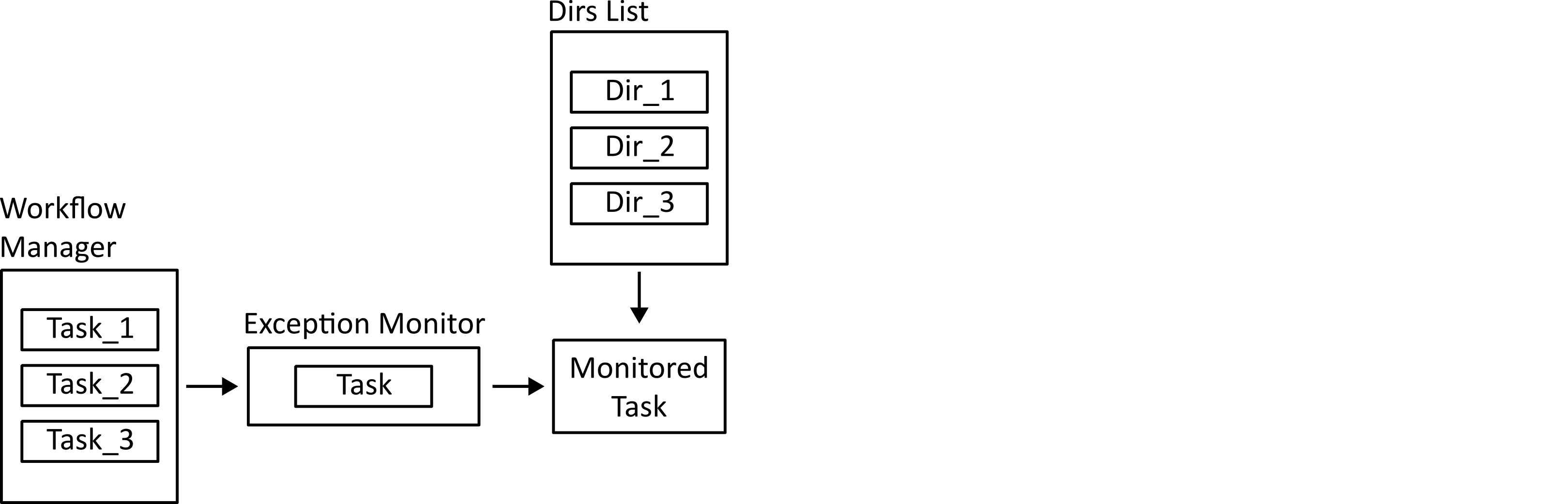
Data provenance
Another challenge I wanted this project to help with in running workflows is managing data provenance. The challenge is to be able to recall exactly how a certain output was produced when it is the product of a complex chain of steps with multiple changeable parameters. It’s important to know this if you want to draw any conclusions or create repeatable processes, but can be quite difficult to manage, especially with code under active development, or if multiple researchers are using data derived from an original dataset on their own differing projects.
The way this project tackles the issue is by using a combination of logging and the python package setuptools-scm. Whenever a workflow is run, a log file is created recording the date/time, the workflow configuration file, and the names of any output files. Along with this, the setuptools-scm package is used to record the version hash from the git version control system (and whether the working directory has any changes since then). This provides enough information to recover the exact code used to create any output file.
Conclusions
Much of this project grew organically from individual processes that I required for my work, along with principles I wanted to demonstrate for others in my research group, such as how to work with configuration files or how to search through directory trees.
Overall, I found that working on this project helped me develop a deeper understanding of the challenges involved in workflow orchestration. In the future I would still probably use an existing tool like Airflow for workflow orchestration, but with a better understanding of how and why those tools work. For example I learned how configuration files need to be structured to contain the right information, how to develop standard interfaces with sufficient flexibility, and what an MVP for data provenance looks like. At the same time, it was working with the users of the software (in my case, when sharing parts of this project with my research group), developing the tools they needed and helping them learn to use them that was one of the most rewarding aspects of the project.